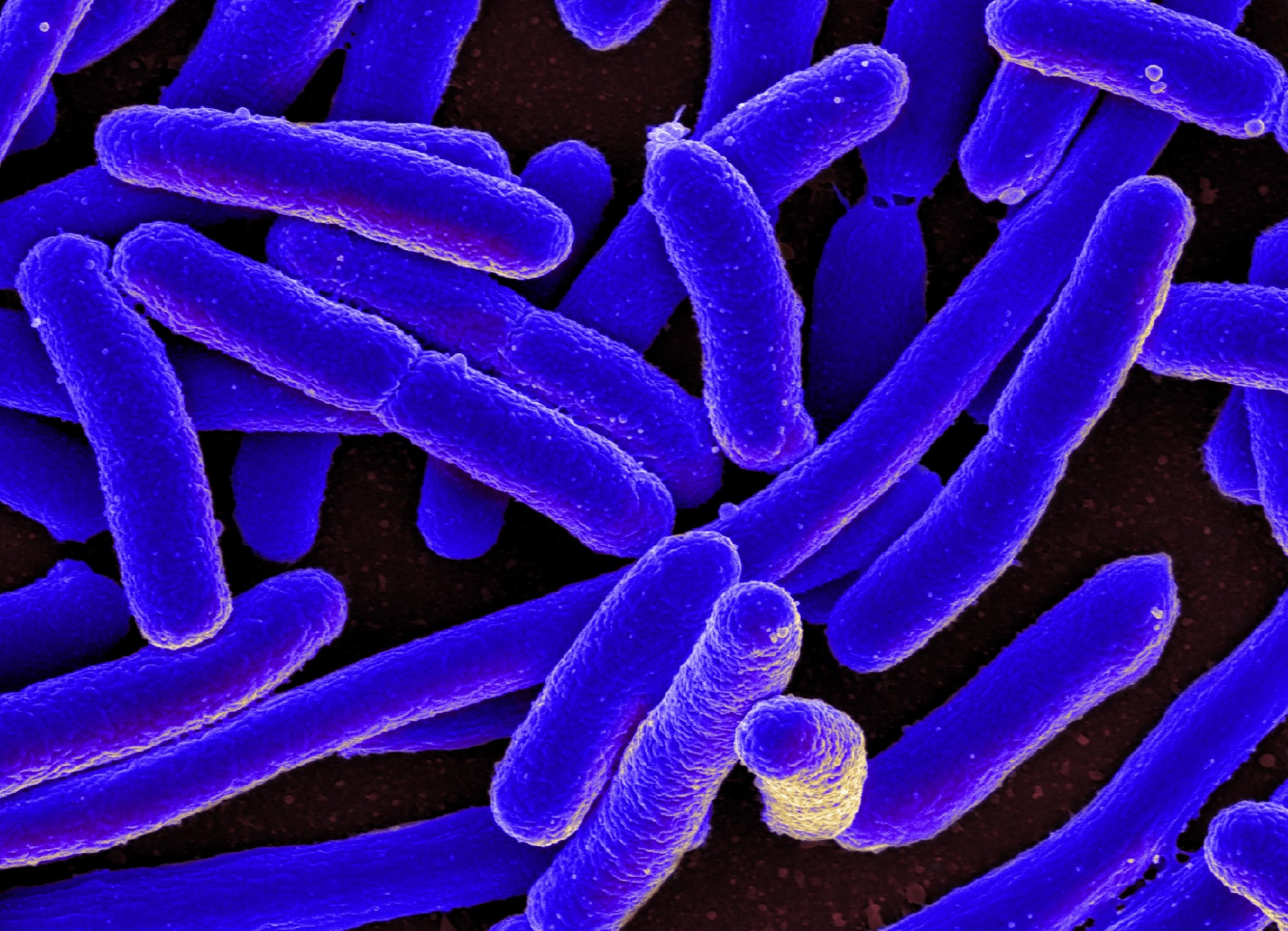
A team of Russian scientists led by Skoltech Professor Mikhail Gelfand has discovered a new process by which the bacteria E. Coli (Escherichia Coli) processes lactose, thus enabling its survival.
Most bacteria are able to survive under a broad variety of ecological circumstances. Often, they are able to adapt their nutritional intake to their surroundings.
The well-known bacteria E. Coli traditionally live in mammalian intestines. But if an E. Coli bacterium is moved to a new habitat, it can typically feed on fatty acids or carbon disulfides, and ultimately other nutrients it can conveniently ingest.
E. Coli most commonly feed on glucose, which tend to enter mammalian intestines following the decomposition of complex carbohydrates. They can also feed on lactose, which is found in dairy products.
In the 1950s, French scientists Francois Jacob and Jacques Monod grappled with the topic of how E. Coli are able to process large amounts of lactose when it’s available, and how they can turn off the enzyme-synthesis mechanism required for lactose utilization when supplies run out. This had long puzzled researchers as idle resources are not evolutionarily beneficial for any organism.
They discovered that the genes located in the chromosomes of E. Coli were organized into unique structures called operons, which enable these bacteria to shut off unnecessary genetic activity, thereby transitioning between the glucose and lactose metabolic mechanisms.
Since then, it has been broadly believed that E. Coli only had one means of utilizing lactose, and that when this genetic activity was shut off, these bacteria would not be able to survive on lactose.
Three years ago, Skoltech scientists teamed up with their counterparts from the Russian Academy of Sciences’ (RAS) Institute for Information Transmission Problems, as well as children from the Moscow-based Summer School of Theoretical Molecular Biology, to study Enterobacteriaceae, the family to which E. Coli belongs.
In the course of their research, they discovered an unusual similarity between the group of Enterobacteriaceae genes that are responsible for processing carbon disulfides, and the combination of genes that other bacterial group- bacilli –use to break down lactose. All these findings came about as a result of bioinformatical techniques.
The RAS Institute of Cell Biophysics joined the group for subsequent experimental research, during which they discovered that E. Coli has an alternative means of processing lactose. In particular, the genes used by these bacteria to process carbon disulfides can also be used to process lactose.
The scientists then switched off the mechanism by which the E. Coli typically processes lactose, and the bacteria continued to grow and multiply on lactose using the newly discovered pathway exclusively.
In a statement, the team said: “These results suggest the possibility that enzymes previously believed to have been highly specialized are actually multifunctional. It also raises numerous additional questions related to biochemical characteristics, specificity, and the affinity to all possible substrates.
“Furthermore, our discovery shows how powerful the integration of bioinformatics and experimental methods can be with respect to solving molecular biological problems,” said Skoltech Prof. Mikhail Gelfand.
The results of the study have been published in Scientific Reports.
Contact information:
Skoltech Communications
+7 (495) 280 14 81